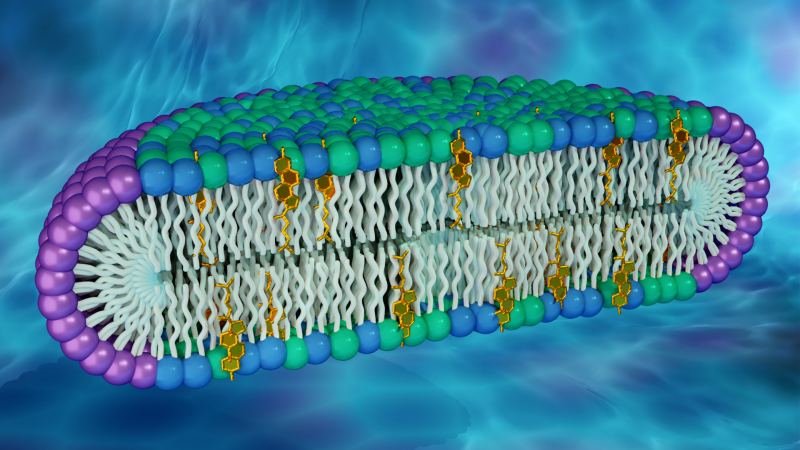
The cell membrane, the wall-like boundary between the cell interior and its outside environment, is primarily made up of two kinds of biomolecules: lipids and proteins. Different lipid species closely pack together to form a double layer, or “bilayer,” the membrane’s fundamental structure, while proteins are embedded within or attached to the bilayer.
Membrane proteins are responsible for various important cellular activities, and their dysfunction can lead to serious health issues. Studying membrane protein structures and how they behave will help scientists better understand their connection to diseases and aid in developing therapeutics.
A team of researchers led by Vanderbilt University has recently shed light on how membrane proteins could be influenced by the lipids around them. By developing a novel type of membrane model, the scientists were able to show that the shape and behavior of a protein can be altered by exposure to different lipid compositions.
The researchers confirmed the artificial membrane’s structure using x-ray and neutron scattering at the Department of Energy’s (DOE’s) Brookhaven (BNL) and Oak Ridge National Laboratories (ORNL). Their findings were published in the Journal of the American Chemical Society.
“This work showed that a protein can change quite profoundly in different membrane lipid environments, and we think this opens up a whole new area of research,” said Charles Sanders, professor of biochemistry at Vanderbilt University and corresponding author of the new study.
Lipid layouts and cell membrane models
Cell membranes are composed of a variety of lipid molecules. Recently, studies have shown that certain lipids in cell membranes can come together to form clusters, also known as rafts. Some scientists suggest that rafts may move throughout the membrane and co-exist with ungrouped molecules. “A lipid raft is like a clique at a party,” said Sanders. “They may move around the party, but it’s always the same people talking to each other.”
His lab is exploring how rafts might impact membrane proteins and cellular activities associated with them. In the new work, Sanders and a team of researchers have created a synthetic membrane capable of incorporating abundant quantities of two lipid molecules thought to make up rafts in cell membranes: cholesterol and sphingomyelin. Their approach involved developing disc-shaped biological structures, known as bicelles, that can produce a simplified model of a cell membrane’s lipid bilayer.
“Cholesterol and sphingomyelin are ubiquitous in cell membranes but have not been present together in previous versions of bicelles,” said John Katsaras, a biophysicist and neutron scattering scientist at ORNL and study co-author. “This new class of bicelles has a lipid composition that we believe is more biologically relevant.”
Complementary techniques give comprehensive analysis
After developing the bicelles, the researchers used small-angle neutron and x-ray scattering techniques to precisely determine the material’s shape and structural organization.
“It’s really hard to confirm the actual morphology of bicelles. Small-angle neutron and small-angle x-ray scattering are the only ways to get a good overall characterization of these particles,” said James Hutchison, a Vanderbilt University researcher and study co-author.
The team used a joint-access program for small-angle neutron and x-ray scattering that allows researchers to more conveniently request beam time at the Bio-SANS instrument at ORNL’s High Flux Isotope Reactor (HFIR) and the Bio-SAXS instrument (LiX) at BNL’s National Synchrotron Light Source II (NSLS-II).
Neutrons can detect light elements like hydrogen, whereas x-rays are more sensitive to heavier elements, which means each scattering technique can reveal unique information about the same material. By using both methods, the researchers built a more accurate model of the membrane system.
“Neutron and x-ray scattering are very complementary to each other,” said Shuo Qian, a neutron scattering scientist at ORNL and study co-author. “Together, those techniques were able to provide a full picture of the bicelle structure.”
Complementary bicelle measurements were also performed using transmission cryo-electron microscopy at Vanderbilt University.
Uncovering new protein properties
To assess how the new model membrane could be used to understand lipid composition and membrane protein relationships, the scientists introduced their bicelles to a well-studied protein fragment, referred to as C99. This fragment makes up one region of a membrane protein called amyloid precursor protein, which experts believe is connected to Alzheimer’s disease.
Using various characterization methods, the team pinpointed differences in the protein fragment’s structure and dynamics when embedded in the new membrane model. Notably, they observed the C99 fragments self-associate with one another in regions that had not previously been reported in other model membranes. The researchers hypothesize these newly uncovered binding sites could play a role in regulating other protein interactions with this fragment.
The team aims to run additional experiments to confirm whether the new bicelle system possesses a lipid raft environment. Scientists have already identified lipid raft properties in artificial vesicles, a spherical hollow biological structure that is enveloped by a lipid bilayer, but not in other small particles, such as bicelles
“There is no known non-vesicle small particle that has lipid raft-like properties,” said Hutchison. “It would be a slam dunk to prove it.”
This work was supported by the National Institutes of Health, the National Science Foundation, and DOE’s Office of Biological and Environmental Research and Basic Energy Sciences.
HFIR and NSLS-II are DOE Office of Science User Facilities. UT-Battelle LLC manages ORNL for the DOE Office of Science. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, please visit energy.gov/science.—by Olivia Trani